1 Statement of need
Scientific results are published in the form of hypotheses, axioms and equations as well as text and diagrams. Likewise, research software is being published more and more frequently. The comprehensibility of scientific results is indispensable for scientific discourse and reproducibility. Hypotheses, axioms and equations are usually published in text form and can be referenced accordingly. Software can be made traceable and referencable through the use of version control software. But what about diagrams? A diagram published in a paper is difficult to trace because the (raw) data is usually not available. However, the traceability of diagrams and the data they contain is not only a challenge in publication but also in everyday research. Diagrams are used for visualization and are therefore often produced for interim results. While the researcher continues the research process with investigations, experiments or simulations, volatile but important information like metadata, background information and details of the data processing are lost.
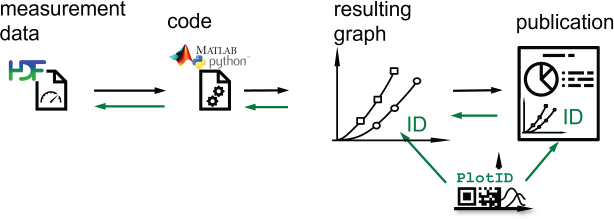
To be able to reconstruct the complete path, a treasure map is needed, starting from a publication, marking major landmarks of the process back to the raw data and metadata. This map needs to be provided along with the product that will be reviewed the most – the created diagram. If diagrams – regardless of whether they are published later or only serve as interim results – are provided with an identifier which connects to previous steps, then traceability can be ensured. Figure 1 shows the order in which crucial elements are created and how the reference chain tracks back.
Research workflow from left to right; afterwards following the chain of references from right to left ‘plotID-referencing’ by Martin Hock, licensed under CC-BY-SA 4.0 
A tool designed to meet these needs must satisfy the following requirements:
Diagrams must have a unique identifier.
The identifier must reference the raw data, relevant metadata and the code used to process the data.
The method must be easy to implement into the existing research workflow.
To reduce the effort of organizing figures along with all necessary data and metadata for later review and reuse, the tool plotID was developed. plotID meets all the above-mentioned requirements. The tool is limited to usage in an existing Python environment, but investigations on enabling independent installation and execution or offering plotID as a web-based service are ongoing. The software depends on multiple other Python libraries. It is currently limited to visualizations from the Matplotlib-library [1] and general picture files such as PNG and JPG.
Researchers often tend to keep an Excel table, noting down manually which data file corresponded with which result along with input parameters. Sometimes an ID system is used (counting up or using the date), but interim results like visualizations – used to verify results – are usually not included. Reviewing and understanding the environment of solutions used in and specifically created for research data management (RDM) remains an ongoing process. The named products in this paragraph are meant to provide some overview and examples but are by no means a comprehensive or rated list. The reviewed solutions range from simple local scripts and libraries (like plotID), backup and synchronization software (for filesystem like ZFS [2] and for folders like rsync [3]), software version control (git [4], svn [5]) and software to extend on version control (git LFS [6], git-fat [7], git-annex [8]) to better handle binary and large data up to dedicated workflow management systems (DataLad [9], DVC [10], signac [11]). Another area of solutions focuses on providing the working environment by integrating documentation with code (Jupyter Notebooks [12]), providing bespoke and versioned Virtual Research Environments (VREs) or offering programmable or fixed – often discipline-specific – schematics in Electronic Laboratory Notebooks (ELNs such as eLabFTW [13], RSpace [14]). With more comprehensive solutions and added functionality for sharing and exporting data, products lean more towards a client-server structure or even a fully hosted product with web and API interfaces. Many solutions are Open Source with Software as a Service (SaaS) offerings. Versioning often uses hashing algorithms for security reasons, thus providing unique identifiers for a specific state (snapshot) out of the box, although those are not always used for identification in user interfaces. Some hosted services implement filesystem-level software to equip each data resource with identifiers to track them independently of their current storage location (iRODs [15]). Structuring and organizing data is part of most RDM solutions and even rather strict ELN products offer to append files, images and comments to their organizational units (a probe or process). Export and sharing of research data along with its metadata is an integral part of most RDM solutions, and most offer more refined features and compatibility than plotID. DVC (Data Version Control) can create plots and visualizations as part of the versioning workflow as well as overlaying multiple versions to show differences between plotted results [16].
While the organization of data, metadata and code as much as identification, versioning and export could be found in several products, the unique feature of applying an ID visibly to a visual representation is not provided in any examined solution. With the big difference in scope, plotID could be implemented as part of a workflow complementing most of the above-mentioned solutions. Only some of the most restrictive ELNs or filesystem-level operations are unlikely to be compatible.
2 Methodology
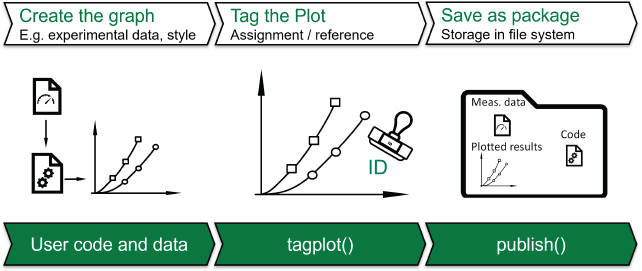
The developed tool plotID is a software solution that covers the needs specified in section 1. The underlying concepts and methods of the software are independent of the programming language. The software aims to support the research workflow shown in Figure 2 and to enable traceability. plotID aims to help during the early research process to decrease the work of making publications reproducible later. To ensure ease of use, the tool has been designed to be integrated seamlessly into existing scripts. For this purpose, a graphical user interface (GUI) has been omitted. Instead, two main functions (building blocks) are provided, which can be inserted into existing user scripts as one-liners. They are the core of plotID. The first module creates a (unique) ID and stamps this ID onto an object containing a visualization, while the second module helps organize all relevant code, software, and data that went into creating this graphic, into one complete package. Furthermore, connectivity to existing identifiers is ensured. If a specific visualization is later chosen to be included in a publication, the ID can be replaced by a permanent identifier like a DOI and the package of code, software and data can be published at the location referenced by the DOI. The ID in the published paper will then directly reference the data, software and code used to create it, thus promoting reproducibility. In the following, plotID is presented in more detail using the Python implementation.
Workflow integrating the plotID core functions ‘plotID-workflow’ by Martin Hock, licensed under CC-BY-SA 4.0 
3 Python – Implementation
The first version of plotID was implemented in MATLAB since this is the most widely used programming language in the local working environment and the language the authors had the highest familiarity with. After reaching a usable state, the focus shifted to rewriting the tool in Python, the second most used programming language (locally). The goal was to make plotID accessible to a broader audience. Globally, Python is a lot more popular than MATLAB with a currently 15 times higher rating on the TIOBE index [17]. Moreover, in contrast to MATLAB, Python better fulfils the requirements for reusable software in the sense of the FAIR1 principles as defined by the Force11 group [18], described for software specifically by Chue Hong et al [19]. Although MATLAB code can be (and often is) Open Source as well this proprietary, commercial software needs to be paid for. This diminishes its accessibility even if older versions are archived properly and provided by the company. In addition to being widely used in the engineering and research community, Python is non-proprietary, Open Source, easy to install and even shipped along many operating systems. Python also offers a package index (PyPI [20] and installer (pip [21]) for easy distribution of software packages.
4 Core functions
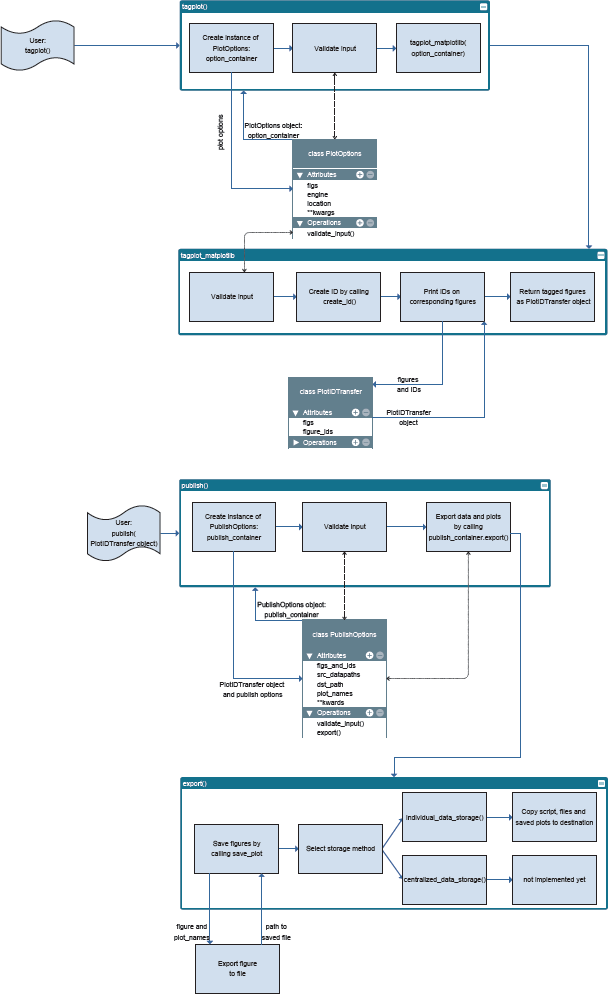
The core functions of plotID are tagplot() and publish(). tagplot() generates an ID and adds this ID to the figure object creating a new container object. publish() takes this container object to bundle the figure, the script file, which plotID was called from and the processed data files and stores them together in a folder named with the ID. In addition, the script is parsed and a list of required dependencies is generated and added as a text file compatible with the Python package installer [21]. New features might bring additional steps with the further development of plotID and a widening of its scope.
System architecture diagram ‘plotID-system-architecture’ by Hannes Mayr, licensed under CC-BY-SA 4.0 
4.1 tagplot()
The tagplot() function creates an ID and tags the figure object with this ID.
4.1.1 ID
tagplot() creates a unique ID (unique in a local system), that consists of a static prefix and a generated part. The prefix parameter is meant to be used to identify a project or organizational unit to which the figure is assigned. The generated part is by default created from the UNIX timestamp in hexadecimal form. As an alternative option, a random number generator can be used. As an alternative to generated IDs, full IDs can be input as a keyword argument, replacing the generated ID. The implementation of the ID is modular, easing the integration of individual needs or sources for IDs. Optionally the ID can be encoded into a QR code for improved machine readability.
4.1.2 Tagging
In Python, there are multiple available packages that can produce visualizations from data. Adding an ID needs to be implemented for many of these engines separately. For now, plotID supports figures created with Matplotlib and raw image files. The ID is added as an attribute to the Python object and the graphical, visible item.
4.1.3 Arguments
Necessary input arguments for tagplot(figs, engine [, **kwargs]):
figs: the figure object or a list of objects, that is to be tagged
engine: the plot/image engine to be used (currently only ‘matplotlib’ and ‘image’ (for plain image files) are supported)
Important optional input arguments are:
figure_ids: IDs that will be printed on the plot. If empty, IDs will be generated for each plot. If this option is used, an ID for each plot has to be specified. Default: empty list. Type: list of strings.
location: Location for ID to be displayed on the plot. The default is ‘east’. Type: string.
rotation: Rotation of the printed ID in degree. Overwrites the value defined by location. Type: float or int.
position: Position of the ID given as (x,y). x and y are relative coordinates with respect to the figure size and must be in the interval [0, 1]. Overwrites the value defined by location. Type: tuple of float.
prefix: Will be added as prefix to the ID. Type: string.
id_method: Creating an ID by Unix time is referenced as ‘time’, creating a random ID with id_method = ‘random’. The default is ‘time’. Type: str, optional
qrcode: Encode the ID in a QR code on the exported plot. Default: False. Type: boolean.
id_on_plot: Print ID on the plot. Default: True. Type: boolean.
There are additional arguments for custom positioning and font of the ID and adjusting the size and position of the QR code.
The function’s output is a PlotIDTransfer Object) which provides a compatible method to transfer the output of all plot engines with additional information, most importantly the ID.
At this point the figure object inside can still be modified, for example, to adjust colours and positioning or to recreate the full plot before exporting a final version.
4.2 publish()
This function starts the export process. The source files of the processed data, the visualization (including the tagged ID), and the script hosting the call to the publish function are copied together into a destination folder.
4.2.1 Script
A function in Python has access to the file path of the script which it was called from. With this, the code for calculations can easily be collected. For this reason, publish() cannot be called from the command line or from within a script that has been started with the ‘python -m’ flag.
For dependent packages, the python compiler can report imported modules with the installed version. Those are then written into a file named “required_imports.txt”. This file can be directly read by the Python package installer to install the code’s dependencies. The import for the plotID package is removed from this list, and in the script file calls to plotID functions are changed to comments, to avoid unnecessary exports. Furthermore, the user has to take care of the necessary Python version (if restrictions apply) and of including additional function files as data paths, if they have not been imported but are still accessed by the executed script.
4.2.2 Data files
Data files are handed over as a list of file or folder paths. Ideally, the script already manages a list of all files that are read during the execution of the script. It is up to the user to control this. By default, the data files are copied to each exported package. For large data files, the ‘centralized’ flag is intended. It is up to the user to decide which resources to add to the export, by placing them in the list of data paths or not.
By default, the data files are copied into each export. The ‘centralized’ selection is optional. With this, the data is copied to a central folder, relative to the export packages. For further exports, the data files are compared to the ones already present and only copied if new data files are selected. With this, a publication on a data repository could encompass the data files in addition to multiple “satellite” folders containing the specific script, parameters and graphics. For HDF5 files, each package can contain an empty HDF5 file that only contains a link to the “real” central data file. While this has proven to be useful in the MATLAB implementation, the Python version aims to include the ‘centralized’ option in a future release.
Remote files on network drives are handled just like local files and while HTTP(s) URLs currently lead to a FileNotFound error, they will be supported in a future release. It is recommended to not add large data or data available from acknowledged repositories into packages meant for publication. plotID will not support additional data or transfer protocols. The exported script should suffice to reference and showcase the usage of remote data sources. Additional commentary or documentation can be added to the export as a data file.
4.2.3 Arguments
Necessary input arguments for publish(figs_and_ids, src_datapath, dst_path [, **kwargs]) are:
src_datapath: This can be a single or a list of file or folder paths for source data and additional function files. The type is a string or a list of strings.
dst_path: This is the destination folder path. If it does not exist, the folder will be created. The type is a string.
figure: This is a figure object, the exact class depends on the plot engine used. This object will be turned into an image file.
Optional input arguments:
data_storage: Currently only ‘individual’ and ‘centralized’ are available. ‘Individual’ will store all data in each exported package, while ‘centralized’ stores the data files in a central folder separate from the packages containing script and image files. To be implemented. Type: string or file path.
plot_name: This is the name for the graphics objects. The type is a string or list of strings. If a single name is passed for multiple objects, a raising number will be added. If no name is passed, the ID will be used as the file name. Type: string or a list of strings.
5 Example script
The following script shows how plotID is used.
10 # %% Import modules
11 import numpy as np
12 import matplotlib.pyplot as plt
13 from plotid.tagplot import tagplot
14 from plotid.publish import publish
15
16 # %% Set Project ID
17 PROJECT_ID = "MR05_"
18
19 # %% Create sample data
20 x = np.linspace(0, 10, 100)
21 y = np.random.rand(100) + 2
22 y_2 = np.sin(x) + 2
23
24 # %% Create sample figures
25
26 # 1. figure
27 FIG1 = plt.figure()
28 plt.plot(x, y, color="black")
29 plt.plot(x, y_2, color="yellow")
30
31 # 2. figure
32 FIG2 = plt.figure()
33 plt.plot(x, y, color="blue")
34 plt.plot(x, y_2, color="red")
38 # If multiple figures should be tagged, figures must be provided as list.
39 FIGS_AS_LIST = [FIG1, FIG2]
In this part, the plotID modules and those necessary to create figures and images are imported. The variable PROJECT_ID is set to provide the static part of the ID. Random data is used to create two figures with Matplotlib.
42 FIGS_AND_IDS = tagplot(
43 FIGS_AS_LIST,
44 "matplotlib",
45 location="west",
46 id_method="random",
47 prefix = PROJECT_ID,
48 qrcode = True,
49 )
Both Matplotlib objects are tagged with a generated ID. Using default options this call fits into a single line.
54 publish(FIGS_AND_IDS, ["../README.md", "../docs", "../LICENSE"], " data")
Files (README.md and LICENSE) and a folder from the code repository are used in place of research data files. The string “data” is the relative path to the destination folder. This also shows that the workflow does not depend on any kind of file format or pre-organized structures. Any kind of data can be used. If the library used for creating the visualization is not (yet) supported, the resulting image file can still be tagged.
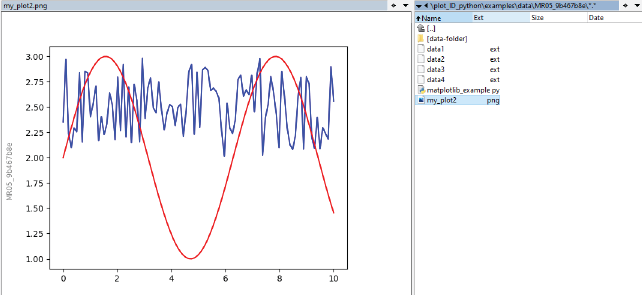
Figure 4 shows the resulting export folder with (renamed) data files, the script file “matplotlib_example.py” and the tagged plot.
Example export folder and tagged plot ‘plotID-example-export’ by Martin Hock, licensed under CC-BY-SA 4.0 
6 Distribution
Providing easy ways to acquire and use the software is important for adoption. The code is Open Source under the Apache-v2.0 license. plotID requires a Python version ≥ 3.10 and is OS-independent. The current release version is v0.3.1. Following Semantic Versioning [22], this indicates that the public API is not considered stable yet.
At this time, the following distribution methods are available and described in the repository’s [23] README file.
6.1 Source Code
The plain source code is publicly available on a GitLab repository located under git.rwth-aachen.de/plotID/plotID_python/ [23] and can be directly downloaded or cloned with git.
1 git clone https://git.rwth-aachen.de/plotid/plotid_python.git
2 cd plotid_python
3 pip install -r requirements.txt
4 pip install .
6.2 Python Package
plotID is listed in the official Python Package Index (PyPI) [20]. The installation is done with the following command:
pip install plotid
Distributing plotID independently from an existing Python installation is one of the aims of later versions. Possible ways to achieve this are providing compiled executable files or a central web-hosted service.
7 Ensuring good software quality
To ensure continuous good software quality, we adhere to best practices and the style guide PEP-8 [24]. This includes comments, docstrings and code formatting. To ensure adherence to these guidelines, automated tests on the code are implemented.
7.1 Unit tests
Python offers various libraries for unit testing. plotID, uses the unittest module [25], which is part of the Python standard library. Tests for each function are defined in the tests folder, along with the runner_test.py script which organizes the execution of the tests, by discovering the test files based on their location. The coverage module measures how much of the code is covered by the tests, and total coverage of less than 95% is considered a failure. The tests are executed by a GitLab CI/CD pipeline [26] with every commit and merge request. Additional Jobs in the pipeline execute Pylint [27] and Flake8 [28] to check against coding style, programming errors and cyclomatic complexity. Commits that fail the pipeline tests cannot be merged into the main branch and will not make it into a release version. In the future, additional tests e.g. against security risks introduced by dependencies and more detailed reports are planned.
7.2 Documentation
To ensure easy access and understanding of the code, Python docstrings [29] have been implemented in the source code from the beginning. The docstrings are compiled into HTML using the Sphinx [30] Python package and GitLab CI-CD [26] creating an automatically generated API reference. The documents are hosted using GitLab Pages [31]. This documentation [32] will be improved by adding the readme, example code, example use cases and an introductory text until version 1.0.
8 Conclusion
The idea of plotID is a simple one: creating snapshots of work. As with many research data management operations, the benefit created through additional effort presents itself only at a later point. Benefits might be harvested by the creators of visualizations themselves by making access to their previous work easier for their own reuse.
The code and open-source implementation are still work-in-progress, but the core functionality is present. There are many ideas to improve and add features reported already and progress in early development happens fast, so many changes should be expected. This paper should be taken as an introduction to the tool and its principles – not as up-to-date documentation. Bug reports, merge requests with code, ideas for features and all feedback are welcome and best voiced in the GitLab repository [23].
Notes
- FAIR: Findable, Accessible, Interoperable, Reusable [^]
9 Acknowledgements
The authors would like to thank the Federal Government and the Heads of Government of the Länder, as well as the Joint Science Conference (GWK), for their funding and support within the framework of the NFDI4Ing consortium. Funded by the German Research Foundation (DFG) – project number 442146713.
10 Roles and contributions
Martin Hock: Conceptualization, Methodology, Coding, Tests, Writing – original draft
Hannes Mayr: Coding, Tests, Methodology
Manuela Richter: Conceptualization, Methodology, Coding
Jan Lemmer: Conceptualization, Methodology
Peter F. Pelz: Project administration, Supervision, Funding Acquisition
References
[1] (2022). “Matplotlib - visualizations with python,” [Online]. Available: https://matplotlib.org/ (visited on 10/10/2022).
[2] O. Rodeh and A. Teperman, “Zfs - a scalable distributed file system using object disks,” in 20th IEEE/11th NASA Goddard Conference on Mass Storage Systems and Technologies, 2003. (MSST 2003). Proceedings., 2003, pp. 207–218. DOI: http://doi.org/10.1109/MASS.2003.1194858.
[3] (2022). “Rsync,” [Online]. Available: https://rsync.samba.org/ (visited on 11/15/2022).
[4] (2022). “Git,” [Online]. Available: https://git-scm.com/ (visited on 11/15/2022).
[5] (2022). “Apache subversion,” [Online]. Available: https://subversion.apache.org/ (visited on 11/15/2022).
[6] (2022). “Git large file storage | an open source git extension for versioning large files,” [Online]. Available: https://git-lfs.github.com/ (visited on 11/15/2022).
[7] (2022). “Jedbrown/git-fat: Simple way to handle fat files without committing them to git, supports synchronization using rsync,” [Online]. Available: https://github.com/jedbrown/git-fat (visited on 11/15/2022).
[8] (2022). “Git-annex,” [Online]. Available: https://git-annex.branchable.com/ (visited on 11/15/2022).
[9] Y. O. Halchenko, K. Meyer, B. Poldrack, D. S. Solanky, A. S. Wagner, J. Gors, D. Mac-Farlane, D. Pustina, V. Sochat, S. S. Ghosh, C. Mönch, C. J. Markiewicz, L. Waite, I. Shlyakhter, A. de la Vega, S. Hayashi, C. O. Häusler, J.-B. Poline, T. Kadelka, K. Skytén, D. Jarecka, D. Kennedy, T. Strauss, M. Cieslak, P. Vavra, H.-I. Ioanas, R. Schneider, M. Pflüger, J. V. Haxby, S. B. Eickhoff, and M. Hanke, “Datalad: Distributed system for joint management of code, data, and their relationship,” Journal of Open Source Software, vol. 6, no. 63, p. 3262, 2021. DOI: http://doi.org/10.21105/joss.03262. [Online]. Available: https://doi.org/10.21105/joss.03262.
[10] (2022). “Data version control – dvc,” [Online]. Available: https://dvc.org/ (visited on 11/15/2022).
[11] (2022). “Signac - simple data management - signac,” [Online]. Available: https://signac.io/ (visited on 11/15/2022).
[12] (2022). “Project jupyter | home,” [Online]. Available: https://jupyter.org/ (visited on 11/15/2022).
[13] N. CARPi, A. Minges, and M. Piel, “Elabftw: An open source laboratory notebook for research labs,” Journal of Open Source Software, vol. 2, no. 12, p. 146, 2017. DOI: http://doi.org/10.21105/joss.00146. [Online]. Available: https://doi.org/10.21105/joss.00146.
[14] (2022). “Rspace eln & inventory,” [Online]. Available: https://www.researchspace.com/ (visited on 11/15/2022).
[15] M. Hedges, A. Hasan, and T. Blanke, “Management and preservation of research data with irods,” in Proceedings of the ACM First Workshop on CyberInfrastructure: Information Management in EScience, ser. CIMS ’07, Lisbon, Portugal: Association for Computing Machinery, 2007, pp. 17–22, ISBN: 9781595938312. DOI: http://doi.org/10.1145/1317353.1317358. [Online]. Available: https://doi.org/10.1145/1317353.1317358.
[16] (2022). “Visualizing plots | data version control - dvc,” [Online]. Available: https://dvc.org/doc/user-guide/experiment-management/visualizing-plots (visited on 11/15/2022).
[17] (2023). “Tiobe index - tiobe,” [Online]. Available: https://www.tiobe.com/tiobe-index/ (visited on 02/24/2023).
[18] M. D. Wilkinson, M. Dumontier, I. J. Aalbersberg, G. Appleton, M. Axton, A. Baak, N. Blomberg, J.-W. Boiten, L. B. Da Silva Santos, P. E. Bourne, J. Bouwman, A. J. Brookes, T. Clark, M. Crosas, I. Dillo, O. Dumon, S. Edmunds, C. T. Evelo, R. Finkers, A. Gonzalez-Beltran, A. J. Gray, P. Groth, C. Goble, J. S. Grethe, J. Heringa, P. A. ’t Hoen, R. Hooft, T. Kuhn, R. Kok, J. Kok, S. J. Lusher, M. E. Martone, A. Mons, A. L. Packer, B. Persson, P. Rocca-Serra, M. Roos, R. van Schaik, S.-A. Sansone, E. Schultes, T. Sengstag, T. Slater, G. Strawn, M. A. Swertz, M. Thompson, J. van der Lei, E. van Mulligen, J. Velterop, A. Waagmeester, P. Wittenburg, K. Wolstencroft, J. Zhao, and B. Mons, “The fair guiding principles for scientific data management and stewardship,” Scientific Data, vol. 3, no. 1, p. 160–018, 2016, ISSN: 2052-4463. DOI: http://doi.org/10.1038/sdata.2016.18.
[19] N. P. Chue Hong, D. S. Katz, M. Barker, A.-L. Lamprecht, C. Martinez, F. E. Psomopoulos, J. Harrow, L. J. Castro, M. Gruenpeter, P. A. Martinez, T. Honeyman, A. Struck, A. Lee, A. Loewe, B. van Werkhoven, C. Jones, D. Garijo, E. Plomp, F. Genova, H. Shanahan, J. Leng, M. Hellström, M. Sandström, M. Sinha, M. Kuzak, P. Herterich, Q. Zhang, S. Islam, S.-A. Sansone, T. Pollard, U. D. Atmojo, A. Williams, A. Czerniak, A. Niehues, A. C. Fouilloux, B. Desinghu, C. Goble, C. Richard, C. Gray, C. Erdmann, D. Nüst, D. Tartarini, E. Ranguelova, H. Anzt, I. Todorov, J. McNally, J. Moldon, J. Burnett, J. Garrido-Sánchez, K. Belhajjame, L. Sesink, L. Hwang, M. R. Tovani-Palone, M. D. Wilkinson, M. Servillat, M. Liffers, M. Fox, N. Miljković, N. Lynch, P. Martinez Lavanchy, S. Gesing, S. Stevens, S. Martinez Cuesta, S. Peroni, S. Soiland-Reyes, T. Bakker, T. Rabemanantsoa, V. Sochat, Y. Yehudi, and R. F. WG, FAIR Principles for Research Software (FAIR4RS Principles), version 1.0, May 2022. DOI: http://doi.org/10.15497/RDA00068. [Online]. Available: https://doi.org/10.15497/RDA00068.
[20] PyPI. (2022). “Pypi · the python package index,” [Online]. Available: https://pypi.org/ (visited on 08/19/2022).
[21] T. pip developers. (2023). “Pip · pypi,” [Online]. Available: https://pypi.org/project/pip/ (visited on 03/08/2023).
[22] T. Preston-Werner. (2023). “Semantic versioning 2.0.0,” [Online]. Available: https://semver.org/ (visited on 03/08/2023).
[23] GitLab RWTH Aachen. (2022). “Plotid / plotid_python · gitlab,” [Online]. Available: https://git.rwth-aachen.de/plotid/plotid_python (visited on 08/19/2022).
[24] (2022). “Pep 8 – style guide for python code | peps.python.org,” [Online]. Available: https://peps.python.org/pep-0008/ (visited on 10/11/2022).
[25] (2022). “Unittest — unit testing framework — python 3.10.6 documentation,” [Online]. Available: https://docs.python.org/3/library/unittest.html (visited on 08/29/2022).
[26] (2022). “Gitlab ci/cd | gitlab,” [Online]. Available: https://docs.gitlab.com/ee/ci/ (visited on 08/19/2022).
[27] GitHub. (2022). “Pycqa/pylint: It’s not just a linter that annoys you!” [Online]. Available: https://github.com/PyCQA/pylint (visited on 08/29/2022).
[28] ——, (2022). “Flake8/index.rst at main · pycqa/flake8,” [Online]. Available: https://github.com/PyCQA/flake8 (visited on 08/29/2022).
[29] (2022). “Pep 257 – docstring conventions | peps.python.org,” [Online]. Available: https://peps.python.org/pep-0257/ (visited on 08/29/2022).
[30] (2022). “Welcome — sphinx documentation,” [Online]. Available: https://www.sphinx-doc.org/en/master/ (visited on 08/29/2022).
[31] (2022). “Gitlab pages | gitlab,” [Online]. Available: https://docs.gitlab.com/ee/user/project/pages/ (visited on 08/29/2022).
[32] (2022). “Welcome to plotid’s documentation! — plotid 0.2.3 documentation,” [Online]. Available: https://plotid.pages.rwth-aachen.de/plotid_python/ (visited on 12/21/2022).